GPCR structural biology: from stubborn to stress-free
Recently, we’ve been thinking carefully about how we can work with G‑protein coupled receptors (GPCRs), a large class of membrane protein in human that mediate and propagate extracellular stimuli to elicit a cellular response. They are involved in sight, smell, touch, development and stress response in the human body, to name a few. Their location on the cell surface and critical role mean GPCRs are an important class of protein for pharmacological intervention of human disease, with an estimated 12% of approved drugs in Europe and America targeting them (1).
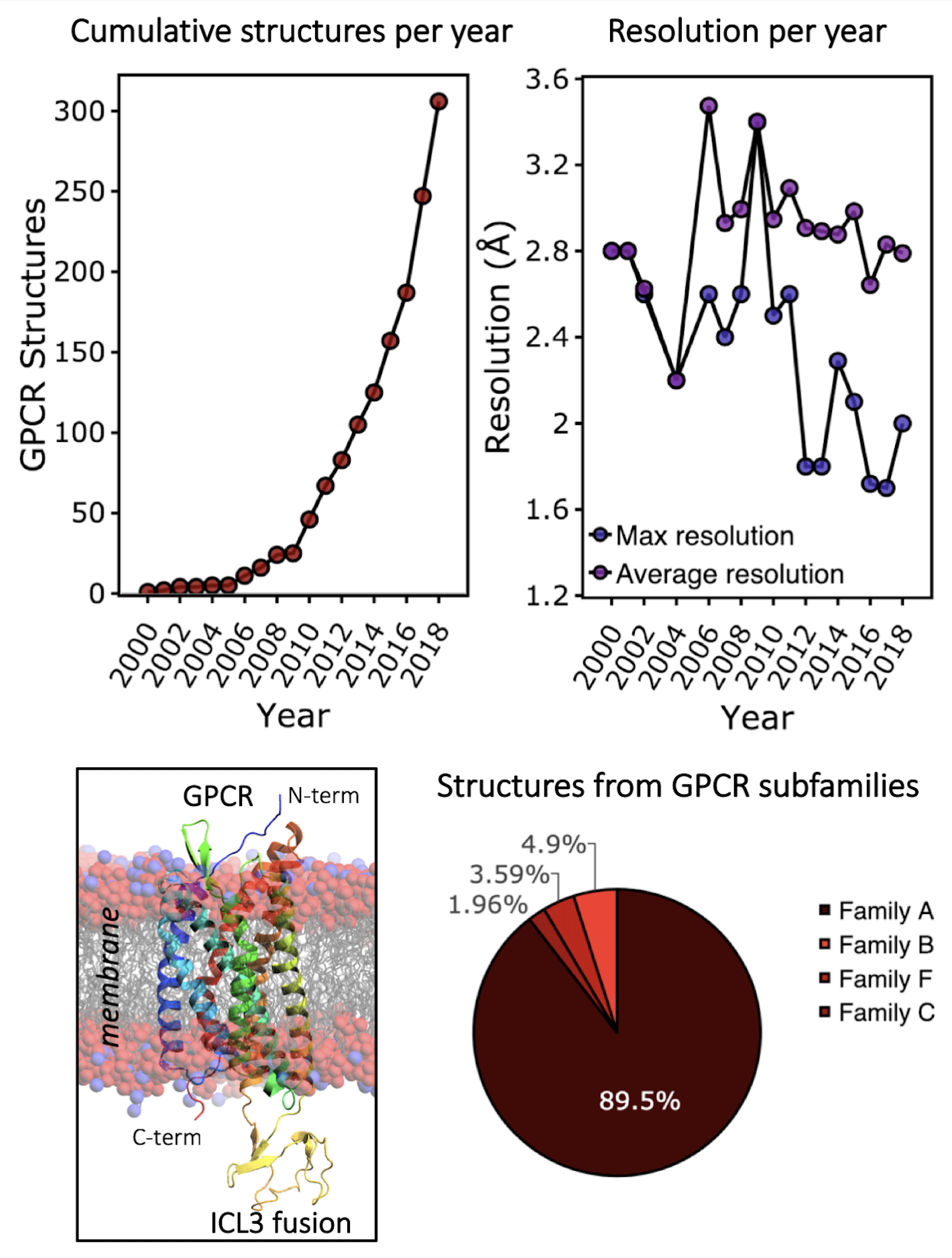
A typical step in the drug discovery process is to understanding where and how small molecules bind their target. This understanding allows rational improvements in chemical properties of the potential drug in order to increase potency and efficacy. Until recently, structural biology of GPCRs was unachievable due to technical challenges related to their biological nature. A number of key advances in technique and understanding between 2004 and 2008 broke this hurdle down, leading to the solution of two key GPCR structures: the β2-adrenergic receptor (2, 3) and the adenosine A2Areceptor (4). Since then, the technical advances that allowed the initial breakthroughs have been transferred to other GPCR family members, leading to an explosion in the number of GPCR structures submitted to the Protein Data Bank (PDB).
So, what was challenging about GPCR structural biology, and what are the steps have become the normal routine for GPCR structure solution?
GPCRs are small alpha-helical membrane proteins located at the plasma membrane. Recombinant expression of human GPCRs must be done in a system that can adequately handle their post-translational processing and modification such as insertion and folding into the membrane, and glycosylation of side-chains. Human embryonic kidney (HEK) cell expression systems have been used, which provide a like-for-like environment for expression. However, the most popular method for GPCR expression is in insect cell systems, with 65% of GPCR structures submitted to the PDB being expressed in Hi5, Sf9or Sf21 cell systems.
A key requirement for successful structure-guided drug discovery is the routine solution of crystal structures to a resolution better than 2.5 Å. There have been some exciting advances in the field of CryoEM that promise to deliver on difficult targets where crystallography has been a bottleneck previously. However, only a handful of GPCR structures have been solved by CryoEM to date, and X-ray crystallography remains the workhorse.
The traditional method for producing protein crystals is the vapour diffusion method, which has been used in the solution of 34% of GPCR structures. However, the Lipidic Cubic Phase (LCP) method for crystallisation of membrane proteins pioneered by the Caffrey Lab has been used in the solution of 65% of GPCR structures (5). LCP is successful because it accommodates membrane proteins in a biphasic, hydrophobic and hydrophilic environment, facilitating crystal nucleation and growth.
A key difficulty with membrane protein work is that membrane proteins must be extracted from their lipid bilayer environment before purification, which is usually done by solubilisation in detergent micelles. Once extracted from the membrane, GPCRs become unstable and prone to unfolding unless specific steps are taken to stabilise them. One way to increase stability is the careful selection of detergent used for GPCR solubilisation. The most popular detergents are decyl maltoside (DM), dodecyl maltoside (DDM), nonyl glucoside (NG) and lauryl maltose neopentyl glycol (LMNG), respectively.
Another method for stabilisation of GPCRs is the introduction of beneficial changes to the amino acid sequence via point mutations. The methods by which these stabilising mutations are found is usually an exhaustive screen of variants by combining iterative rounds of alanine scanning followed by position mutagenesis (6). Two-thirds of all the GPCR structures solved so far have had at least one stabilising mutation introduced.
A final method for stabilising GPCRs is the introduction of a super-stable soluble fusion partner or binding of antibody derivatives or nanobodies to the GPCR. This can reduce conformational flexibility and increase surface area for crystal contact formation, thus increasing crystallisation success and resolution. The most popular fusions are phage T4 lysozyme and thermostabilised apocytochrome b562RIL (BRIL), and the most popular positions for their insertion is in intracellular-loop 3 or at the N-terminus, respectively.
By following these key steps, there is a clear recipe for success in GPCR structural biology, turning a challenging area into a routine science in drug-discovery pipelines.
To find out how Peak Proteins could help you with your membrane protein projects, have a look at our website or contact us directly at info@peakproteins.com



